projects
Collected below are projects for my own PhD research in computational neuroscience and cognitive science, stuff I contribute to, as well as some fun personal projects. Associated code and papers are included wherever possible. You can find all of them on Google Scholar.
dissertation
You can find my full thesis here, and the defense talk is available on YouTube, thanks to the pandemic. This pretty much summarizes most of the works listed below.
topic: decomposing neural field potentials
what: In this set of works, I build computational models of neural field potentials (e.g., ECoG) from populations of neurons, and then try to work backwards and infer variables about the neural population from both simulated and experimental field potential recordings.
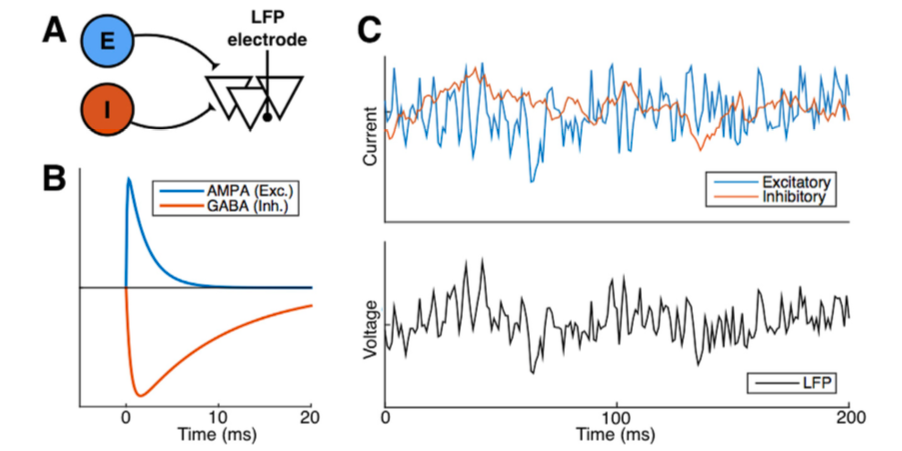
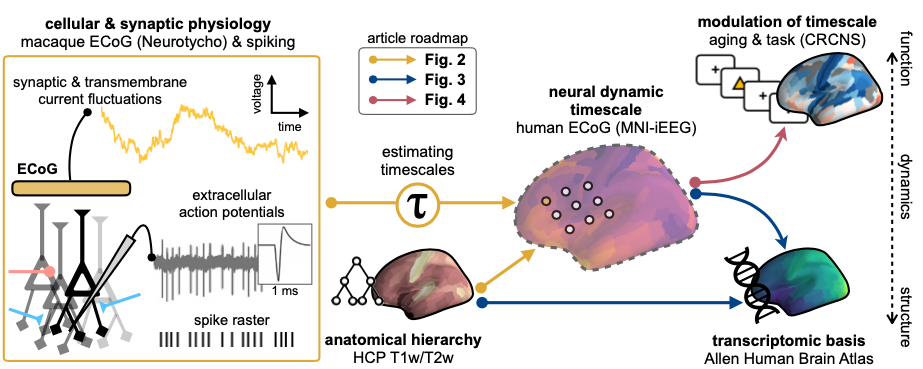
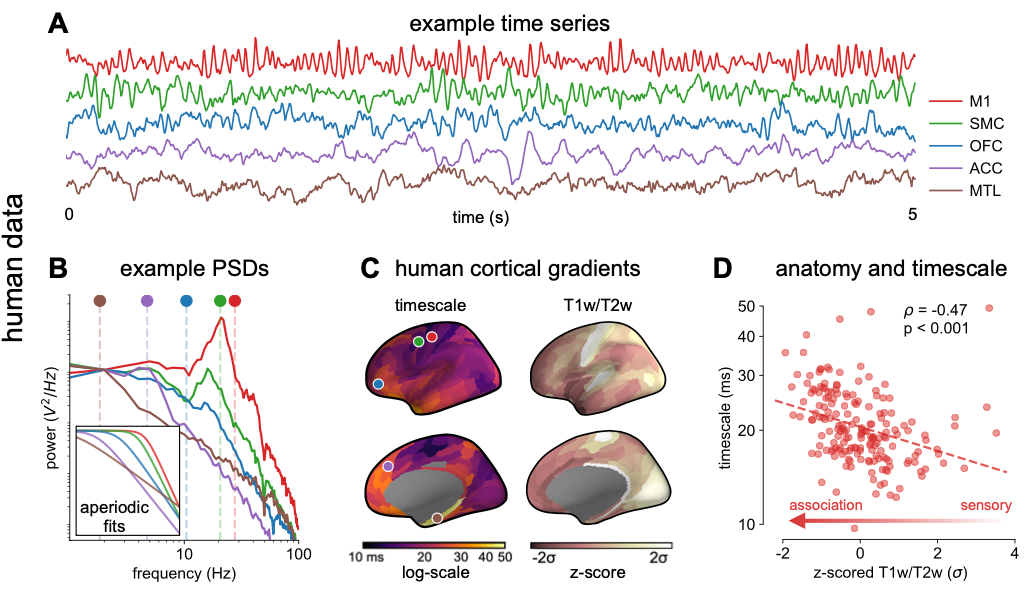
why: Features of the neural field potentials, such as oscillations and asynchronous activity, are routinely used in cognitive neuroscience as biomarkers of behavior, cognition, and disease, yet we have very little idea about how these mesoscale population-level signals come about. Personally, rather than just measuring them, I think having a theory-driven forward model of the signals we use would help us in better linking them with the behavioral variables we’re ultimately interested in.
measuring neuronal timescales across human cortex (2020)
measuring asynchronous (stochastic) population firing (on-going)
inferring synaptic excitation-inhibition balance (2017)
relating spectral power law changes to population firing rate (2016)
App beautiful shiny.io app courtesy of Erik Peterson
topic: development of network activity in brain organoids
what: Collaborating with the Muotri Lab at UCSD, we (read: they) have developed human induced pluripotent stem cell (hiPSC)-derived brain organoids that display spontaneous and oscillatory network activity over a long developmental period. My role has since been to analyze the electrophysiological data (spiking and LFP) to characterize the developmental trajectory of network activity and relating it to structural and molecular variables, as well as unexpectedly learning a lot about electrical activity-dependent neurodevelopment in-vivo.
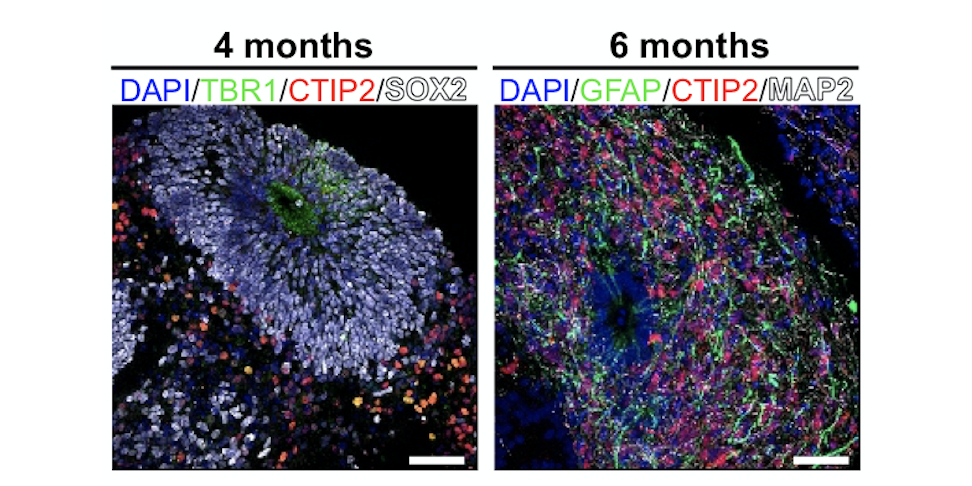
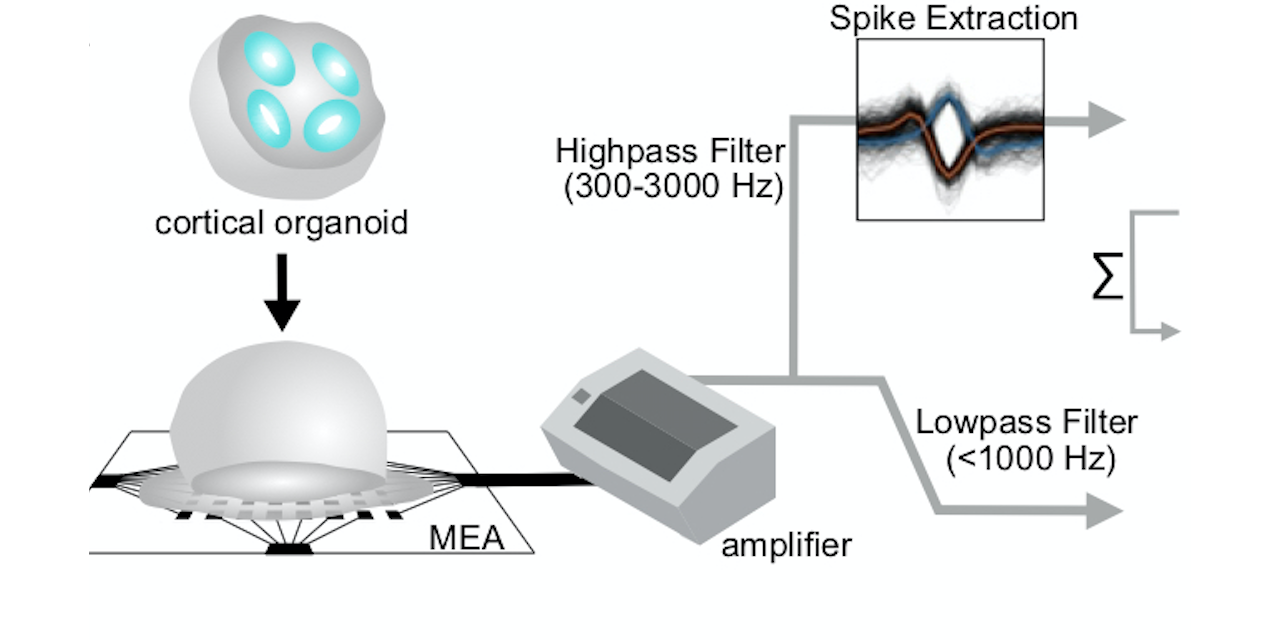
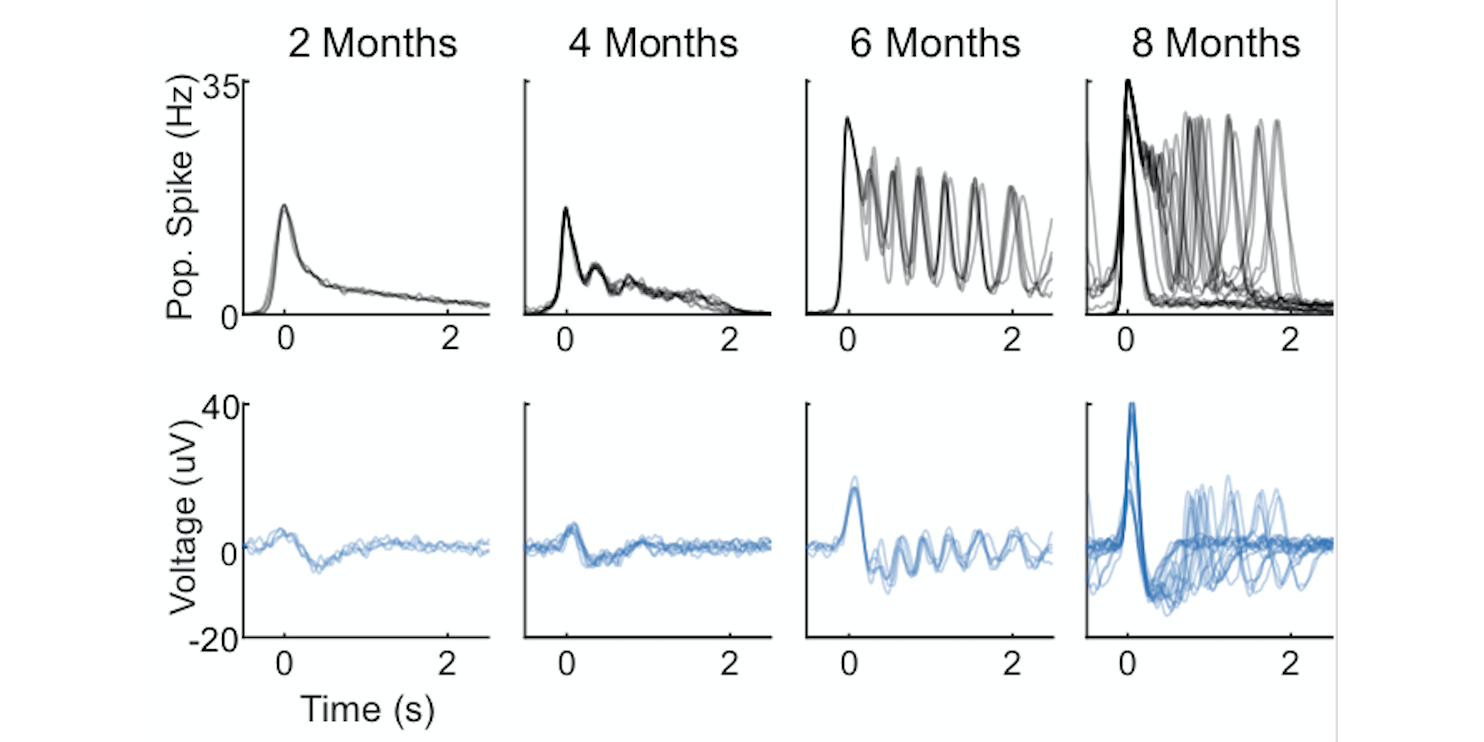
why: Figuring out how neurons in the brain organize themselves over time, both structurally and dynamically, is one of the biggest ongoing challenges in systems and cognitive neuroscience. Brain organoids have recently emerged as a great model for neurodevelopment, having been shown to mimic in-vivo neurodevelopment structurally and in gene expression. By also tracking the emergence of network dynamics over time, as well as continuing to engineer models with higher fidelity, I’m hoping we can dissect how neural circuits organize themselves from the moment neurodevelopment begins, and learn when and how it goes wrong in diseases.
complex oscillatory waves emerging from cortical organoids model early human brain network development
(repo contains code for preprocessing Axion Maestro multi-electrode array data.)
topic: defining “cognition”
what: This hacked-up project that first took a week to implement has turned into a full ideological commitment. I scraped from scientific publications usages of “cognitive processes” words, like ‘attention’ and ‘memory’, and compare 1) which of these cognitive processes are similar, at least from the (crude) perspective of how scientists use them, and 2) how cognitive scientists and neuroscientists use them differently.
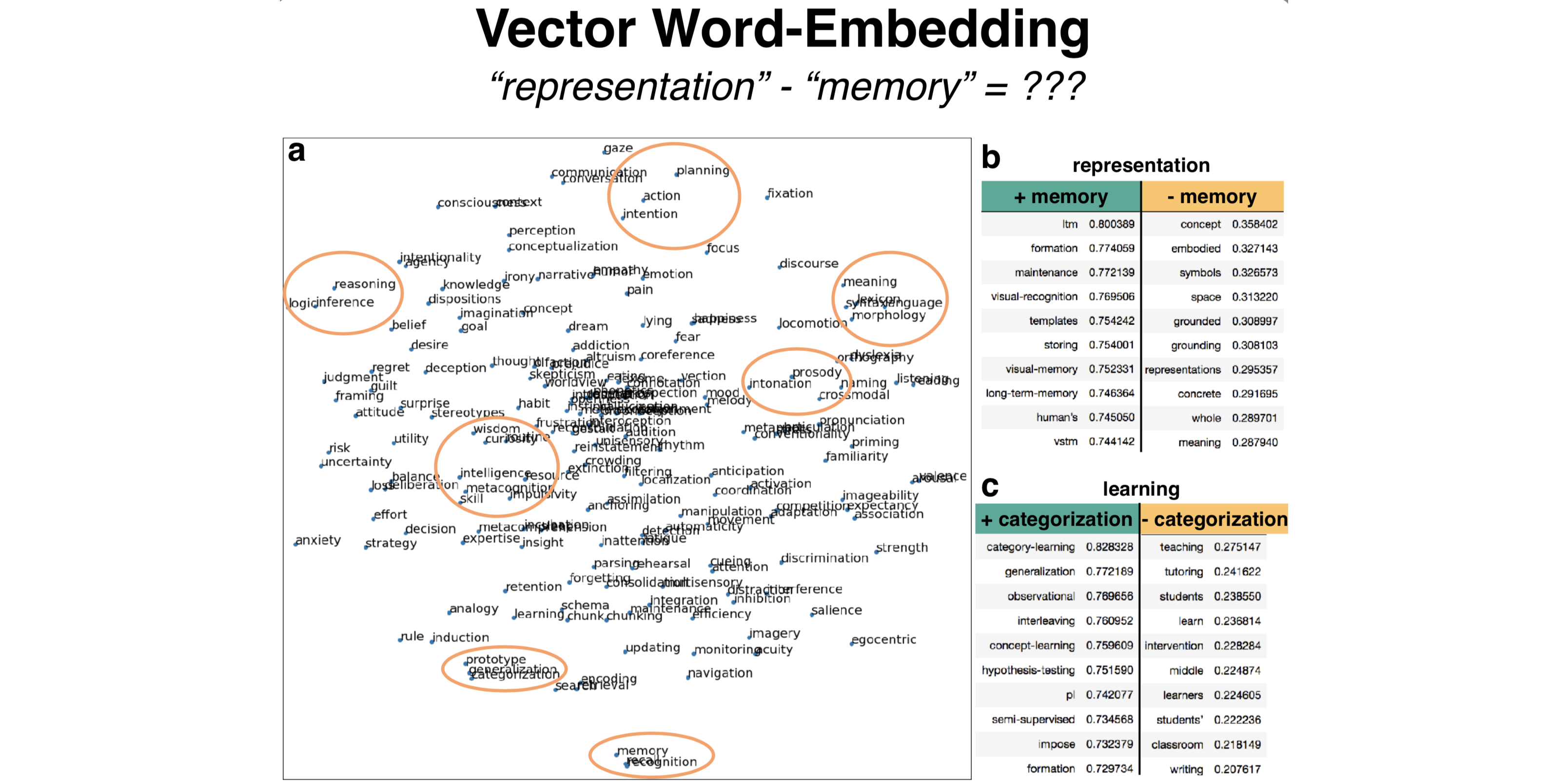
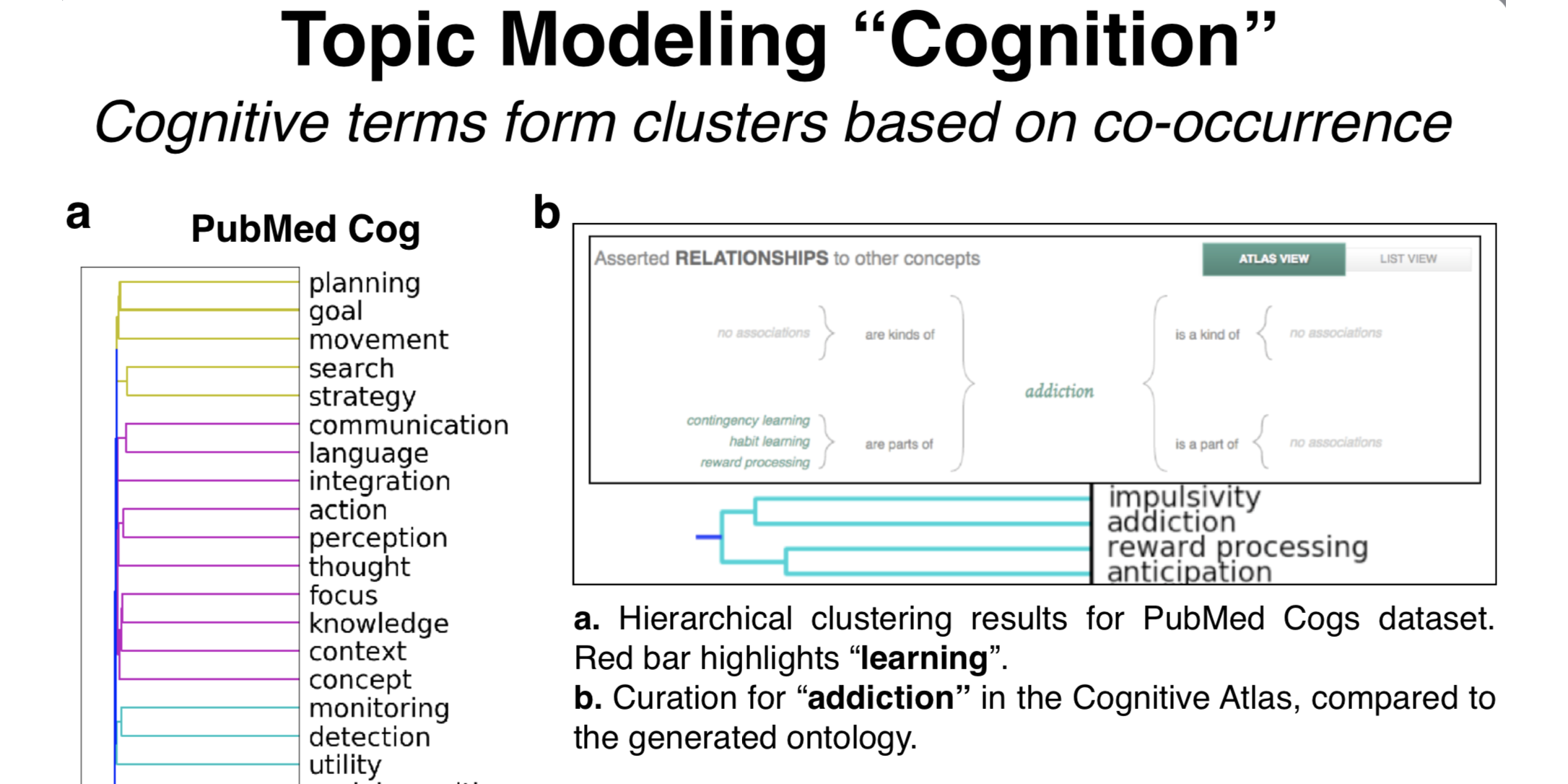

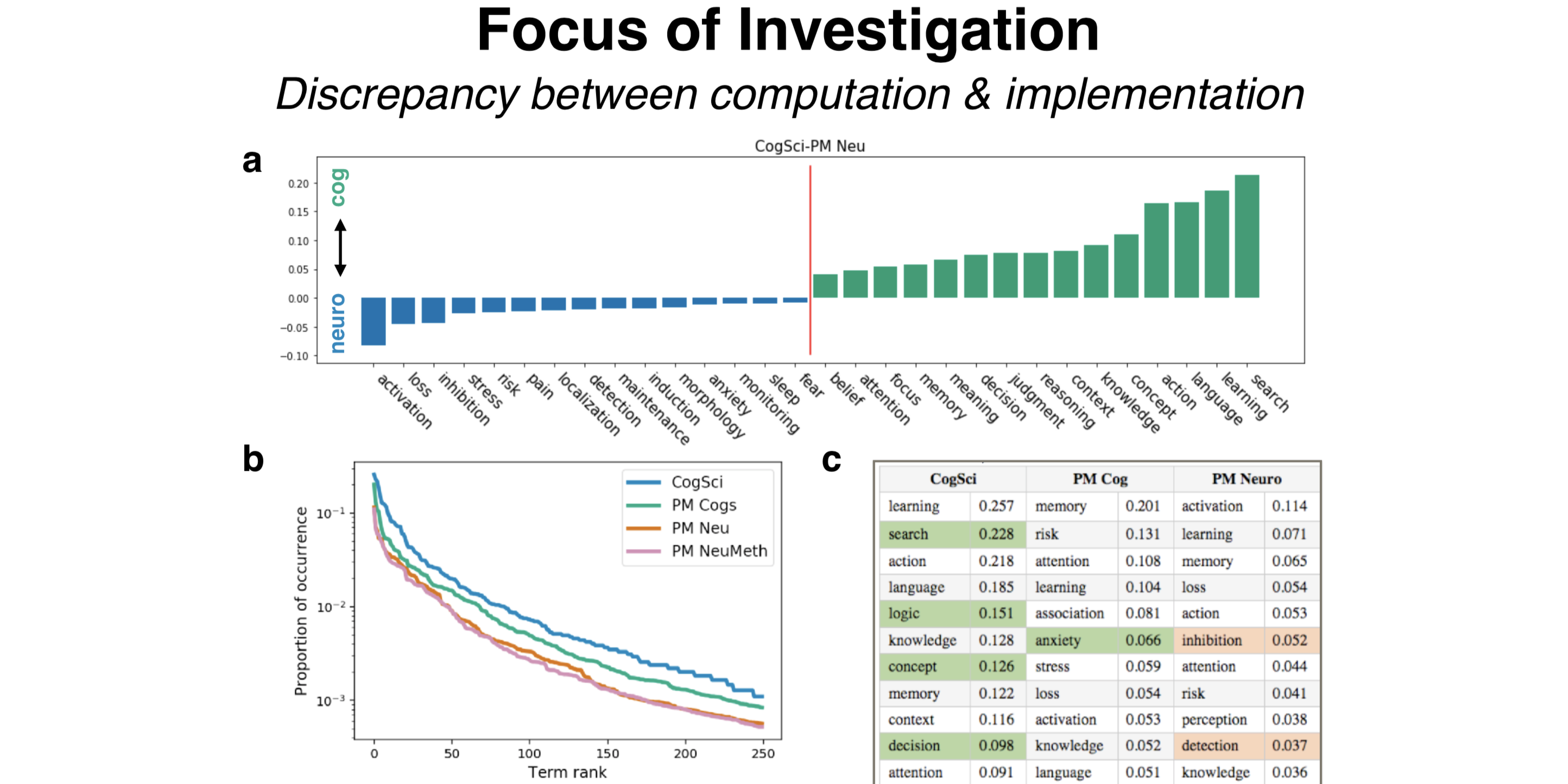
why: Much of cognitive science and cognitive neuroscience aim to characterize and understand “cognitive processes” - processes of the mind that are internal to the organism. Going into my PhD as a freshly minted engineer, it bothered me to no end that there were very few concrete and non-circular definitions for these processes. Precisely because they are internal processes, definitions are hard to pin down, so I took a behaviorist approach and looked at how cognitive scientists and neuroscientists use them in relation to each other in practice - sometimes using the same words to mean very different things - by generating a data-driven ontology scrapped from scientific abstracts. Inspired by Cognitive Atlas, Neurosynth, ERP-SCANR, and BrainSCANR.
